Hierarchical Sampling and Scoring¶
Hierarchical Search¶
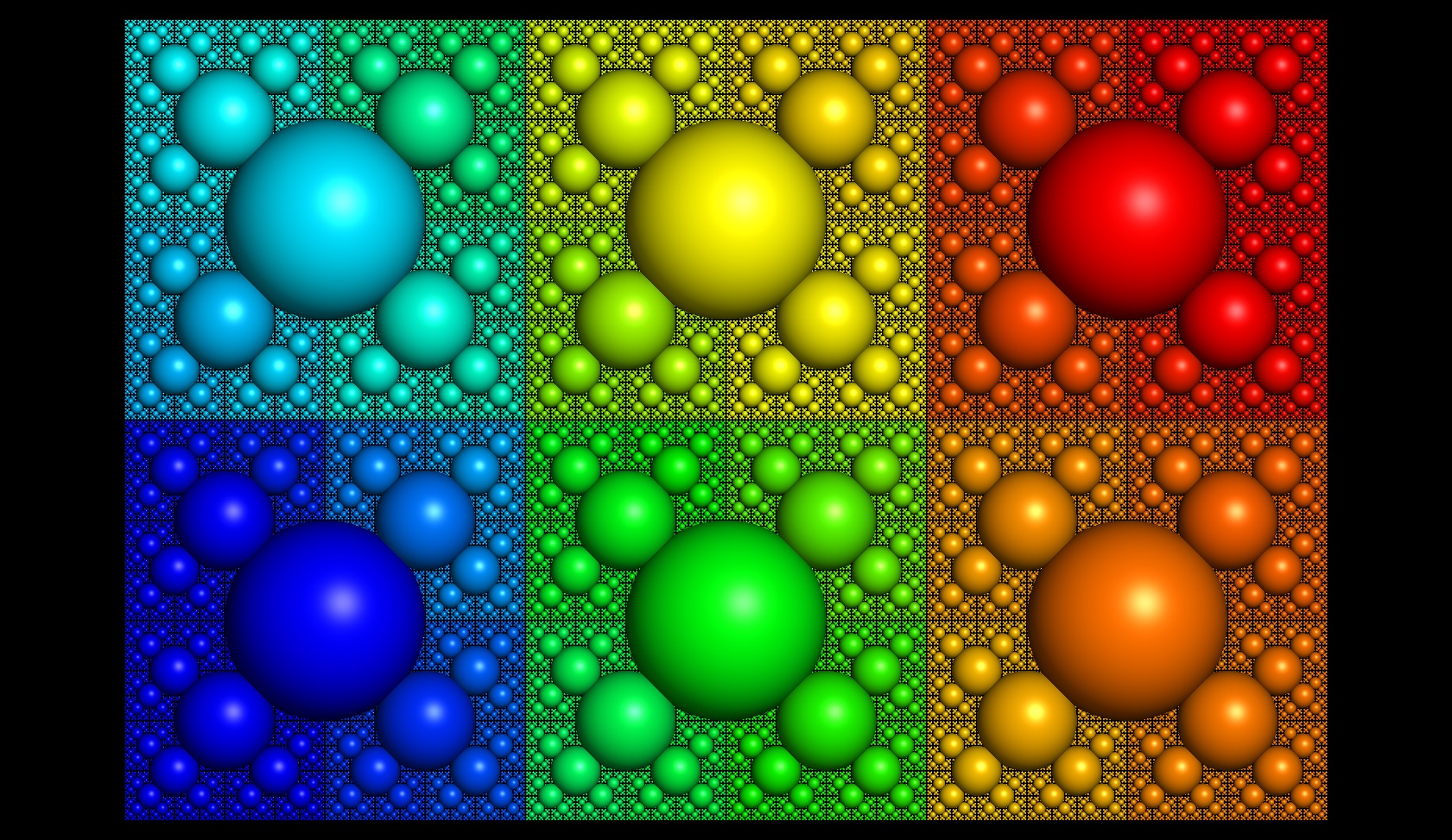
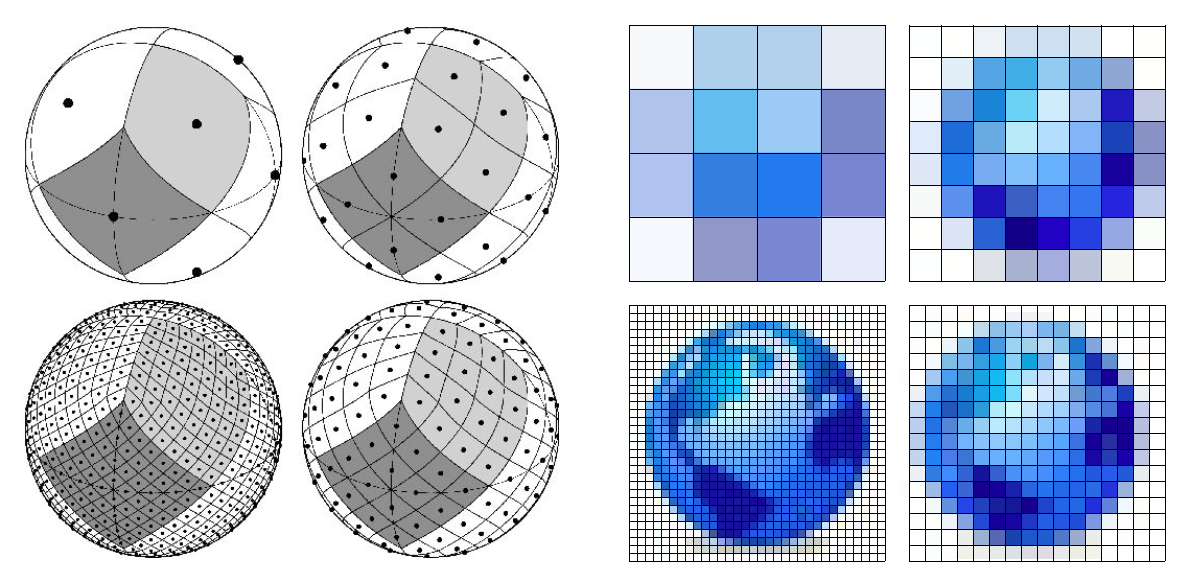
In SCHEME all DOFs are indexed in a nested hierarchy. The number of samples along each dimension doubles with each increase in resolution in a nested manner (see illustrations). This nesting forrest of conformations can be searched efficiently by first examining the lowest resolution points with a correspondingly low resolution objective function, then choosing a best-scoring subset for further analysis. All the children of this subset are then examined with a higher resolution objective function, and another best-scoring subset is generated. This process repeats to the desired precision, probably 5-10 iterations for most protein sampling problems.
Below is an illustration of a nesting hierarchical search grid (from Healpix):
Example:
Sampling Everything¶
Say we search a 6 dimensional DOF space, a simple asymmetric docking problem, and the coarsest reasonable resolution is 4 samples per dimension, or 4,096 points, corresponding to a sample roughly every 8Å. The total number of samples to evaluate grows as the 6th power of the number of samples per dimension:
Number of samples at resolution 8.0Å is 4,096
Number of samples at resolution 4.0Å is 262,144
Number of samples at resolution 2.0Å is 16,777,216
Number of samples at resolution 1.0Å is 1,073,741,824
Number of samples at resolution 0.5Å is 68,719,476,736
Number of samples at resolution 0.25Å is 4,398,046,511,104
Number of samples at resolution 0.125Å is 281,474,976,710,656
Sampling Hierarchically¶
obviously, simple enumerative sampling is impossible at a reasonable resolution, even in this simple case of two body docking. If instead we adopt an iterative hierarchical search in which we prune away areas of the search space which are not likely to contain good solutions, the number of samples which must be evaluated no longer grows exponentially. Say at each stage we keep only the best 10,000 samples, and evaluate all their “children” in the next stage. In this 6 dimensional case, each sample has 64 children so we evaluate at most 640,000 points at each stage:
Number of samples at resolution 8.0Å is 4,096
Number of samples at resolution 4.0Å is 262,144
Number of samples at resolution 2.0Å is 640,000
Number of samples at resolution 1.0Å is 640,000
Number of samples at resolution 0.5Å is 640,000
Number of samples at resolution 0.25Å is 640,000
Number of samples at resolution 0.125Å is 640,000
So the total number of samples is 3,466,240. Scheme scoring is quite fast, on a 4-core cpu in the current implementation, this would take about 30 seconds.
Is Hierarchical Search Really Global Search?¶
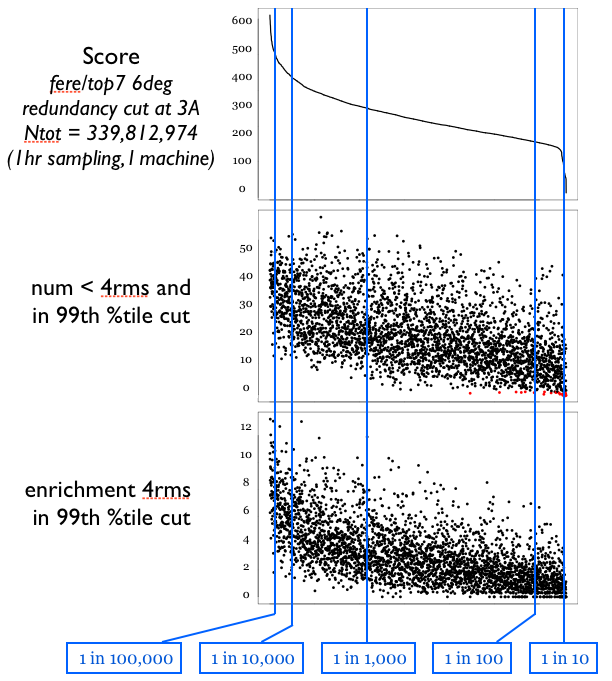
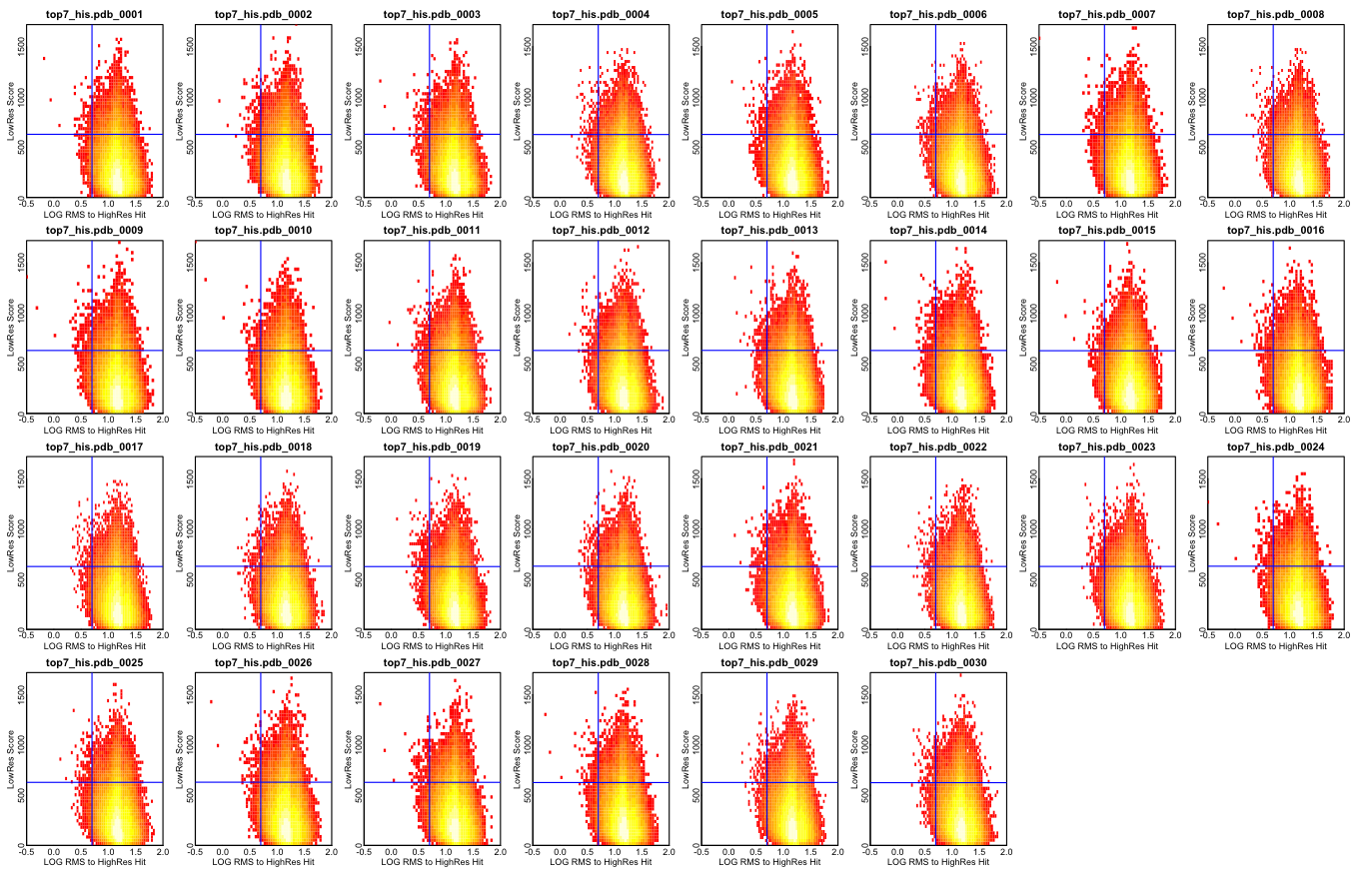
Of course, we may miss some solutions by discarding all but the top 10,000 at each stage. In some simple empirical tests, is seems that we don’t miss too much. The figure below shows a comparison between two sampling runs, one sampling roughly every 0.8Å with ~340 million samples, and one sampling roughly every 4.0Å with ~100,000 samples. The top panel shows a redundancy filtered distribution of scores for the large sampling run. Because we want to examine how “deep” into the global landscape of the larger set we can “see” based on the smaller set, the large set is pruned very aggressively on redundancy, no structures within the set are within 3Å rmsd of each other. The X axis on all three plots corresponds to the sorted index of structures in this high-res, redundancy-filtered reference set. The middle panel plots, for each structure S in the reference set, the number of structures in the low-resolution set which are [1] in the top 1/100th of the set by low-res score, and thus would be candidates for more refined sampling, and [2] are within 4Å rmsd the reference structure S, and thus “cover” S and could recover it in a search. It seems from this data that the hierarchical search approach should not prune away too many “good” candidates.
This test was done with “averaging” type scoring, NOT the superior bounding type scoring.
clash checking¶
We can totally solve the hierarchical clash checking problem using an euclidean distance transform (EDT) on the excluded volume elements (I say we call them “Thorns”). This will give, for each atom, a depth below the surface of the molecule. If you’re clash-checking at a resolution of, say, 2A, you simply set the atomic radius at (depth-2A), ignore if below zero, and your good. Will be easy to use the existing clash-check machinery with this extension. Also gives the opportunity for very fast approximate clash checking by picking some small representative set of atoms with high depths – the depth can be used as a radius for clash-checking, so atoms with high depths can exclude alot of volume with only a handful. Obviously, picking the representatives will take some care, but I think we’ll get alot of speedup out of a fast non-strict clash check even if it isn’t prefect.
Nesting Enumerated State Tree (Nest)¶
All DOFs are managed by Nest objects, which define a hierarchical tree of all allowed states. Nests can control: - Stem transforms in the Bouquet (scene tree) - Rose conformation DOFs (clustered fragments, parametric backbones, etc) - Helical symmetry DOFs (helix and screw axes)
Indexing Info Partially Obsolete!¶
Zorder Indexing¶
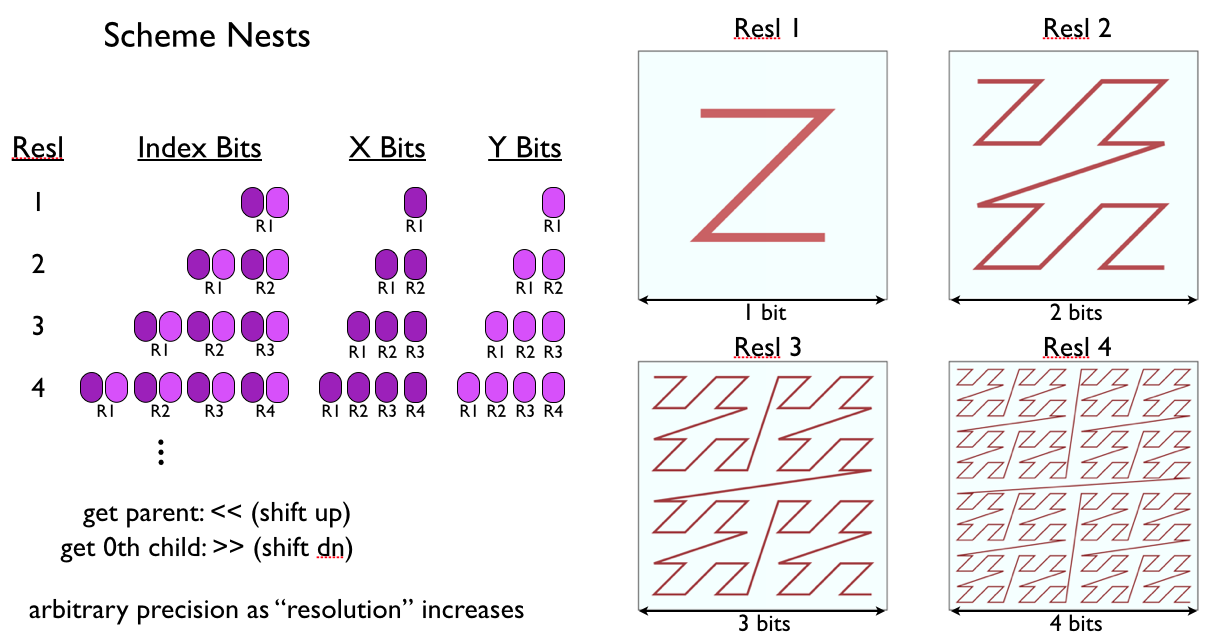
Illustration of basic Nest indexing, which using “z-order” indexing and “morton numbers” in order to (1) make sure siblings in the hierarchy have contiguous indices, avoiding explicit storage and listing, and (2) increase memory coherency by ensuring similar conformations are sampled together. Simple bitshifts are all that’s needed to get the parent or children of an index.
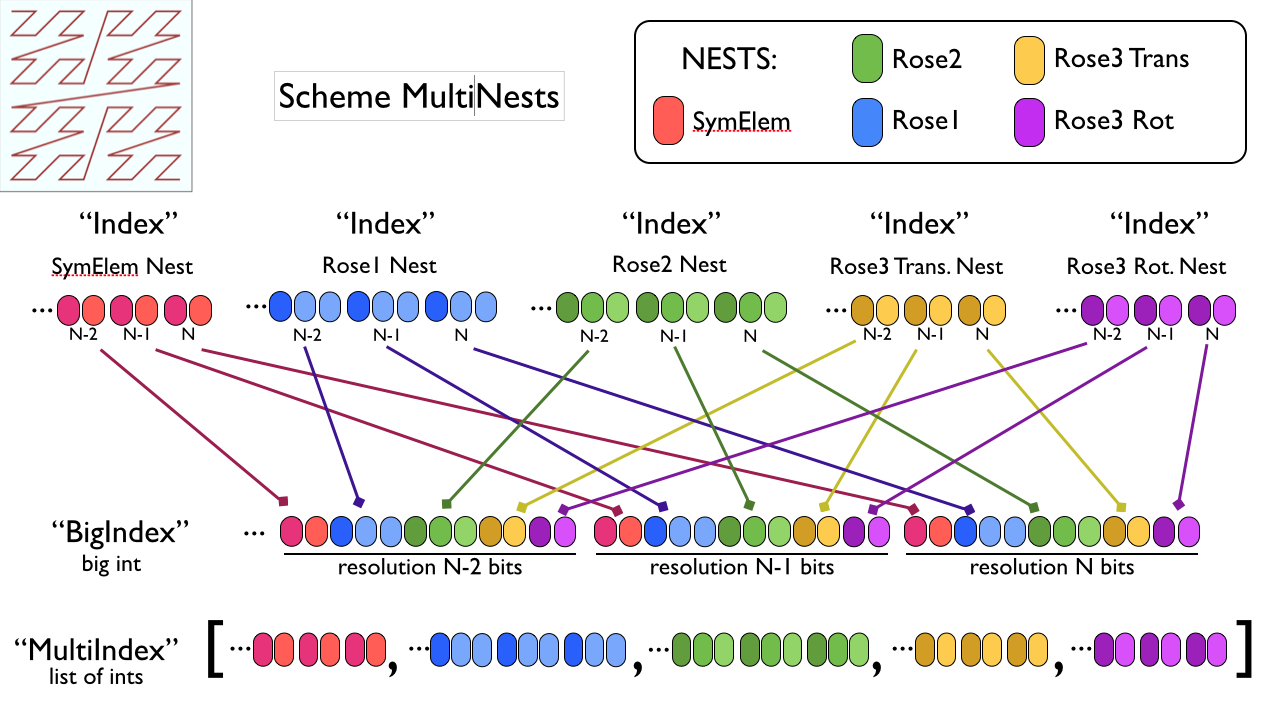
MultiNest Indexing¶
Illustration of a Scheme MultiNest index structure, showing the “BigIndex” strategy which preserves zorder indexing for a set of Nests, and the “MultiIndex” strategy which just lists individual indexes. We may need to employ some mix of these constructs to cope with the indexing headaches involved with Hierarchical Packing.
Indexing Challenges¶
must be able to sort and iterate based by Zorder.
must be able to decompose by subsets of Nests within MultiNest (for pair-decomposition)
must be able to decompose by individual Nests
must be able to efficiently represent a list of indices to be evaluated in a Course of sampling
BigIndex with arbitrary precision implementation is terrible, each on the heap
MultiIndex maybe better, if done with 2D array, but how to increment & sort efficiently
Preferably, list only the “parents” at a given level in the hierarchy, as all children of each parent will be checked
Here’s an illustration of what some of these different indexing types are
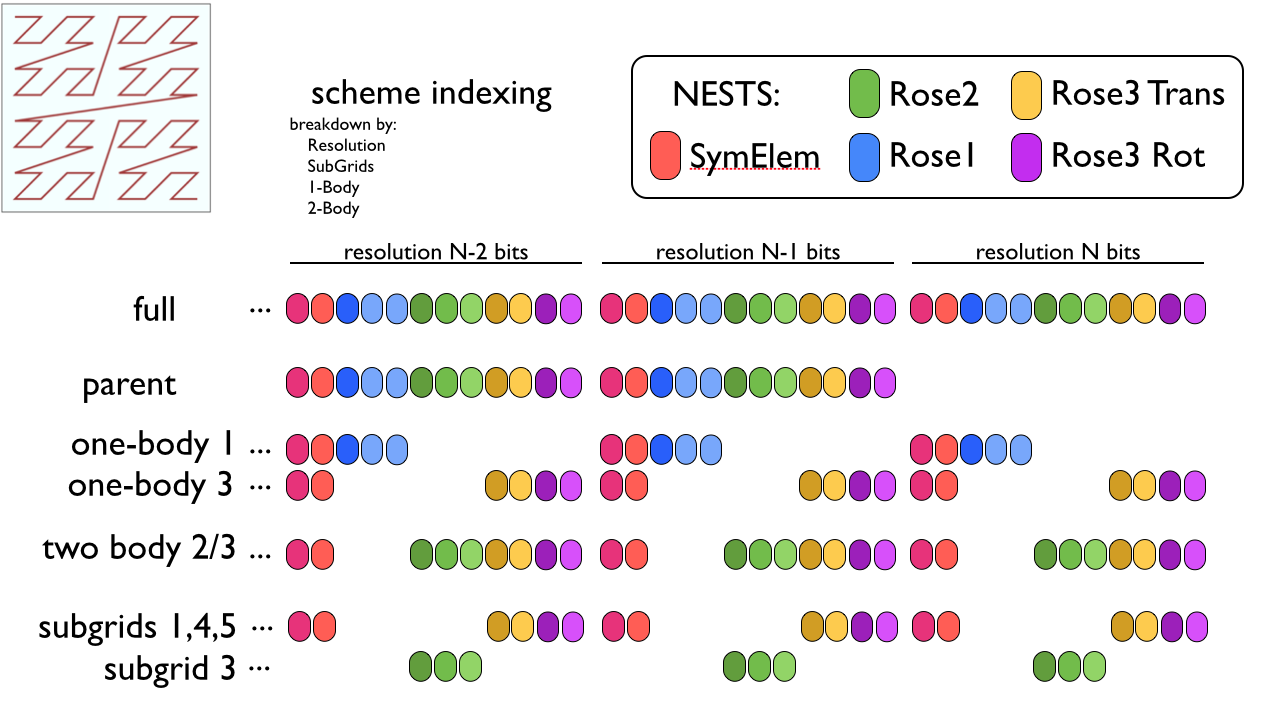
I propose some kind of index manager structure that can assist with all this. It’s implementation can at first be simple and slow, and later be replaced with something more nastily efficient with the same interface. Something like this:
class MultiIndexSet
- -Types
strong_typedef Size IndexMS // sample number in current Course
strong_typedef Size IndexG // Index of Nest (to select a nest)
strong_typedef Size IndexRB // index of Rose
strong_typedef Size Index1B // Index of one-body states
strong_typedef Size Index2B // Index of two-body states
- -Attributes
explicit_parent_indices : BigIndex
- -Operations
IndexMS nstate_total () // total number of samples to be done in set
Index nstate_grid ( IndexG ) // size() of one Nest
Void index_grid ( IndexG, IndexMS ) // Nest index
Index1B nstate_onebody (IndexRB ) // number of distinct one-body states in set for a body
Index1B index_onebody ( IndexRB, IndexMS )
Index2B nstate_twobody ( IndexRB, IndexRB )
Index2B index_twobody ( IndexRB, IndexRB, IndexMS )